Software
Our code is developed under vollgerlab and fiberseq on GitHub.
Tools

Visualization
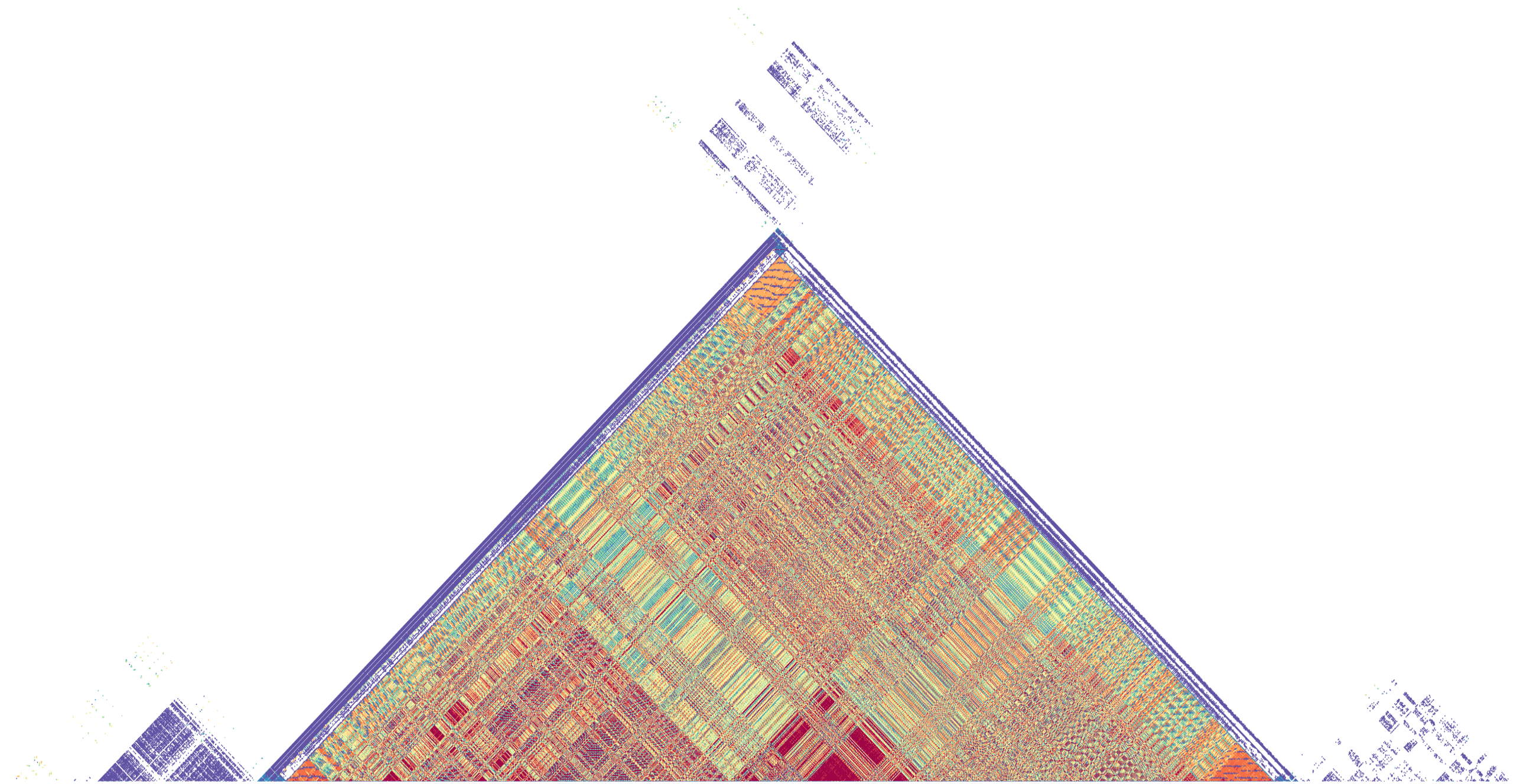

StainedGlass
HTML★ 128
Interactive visualization of tandem repeat structures with identity heatmaps.

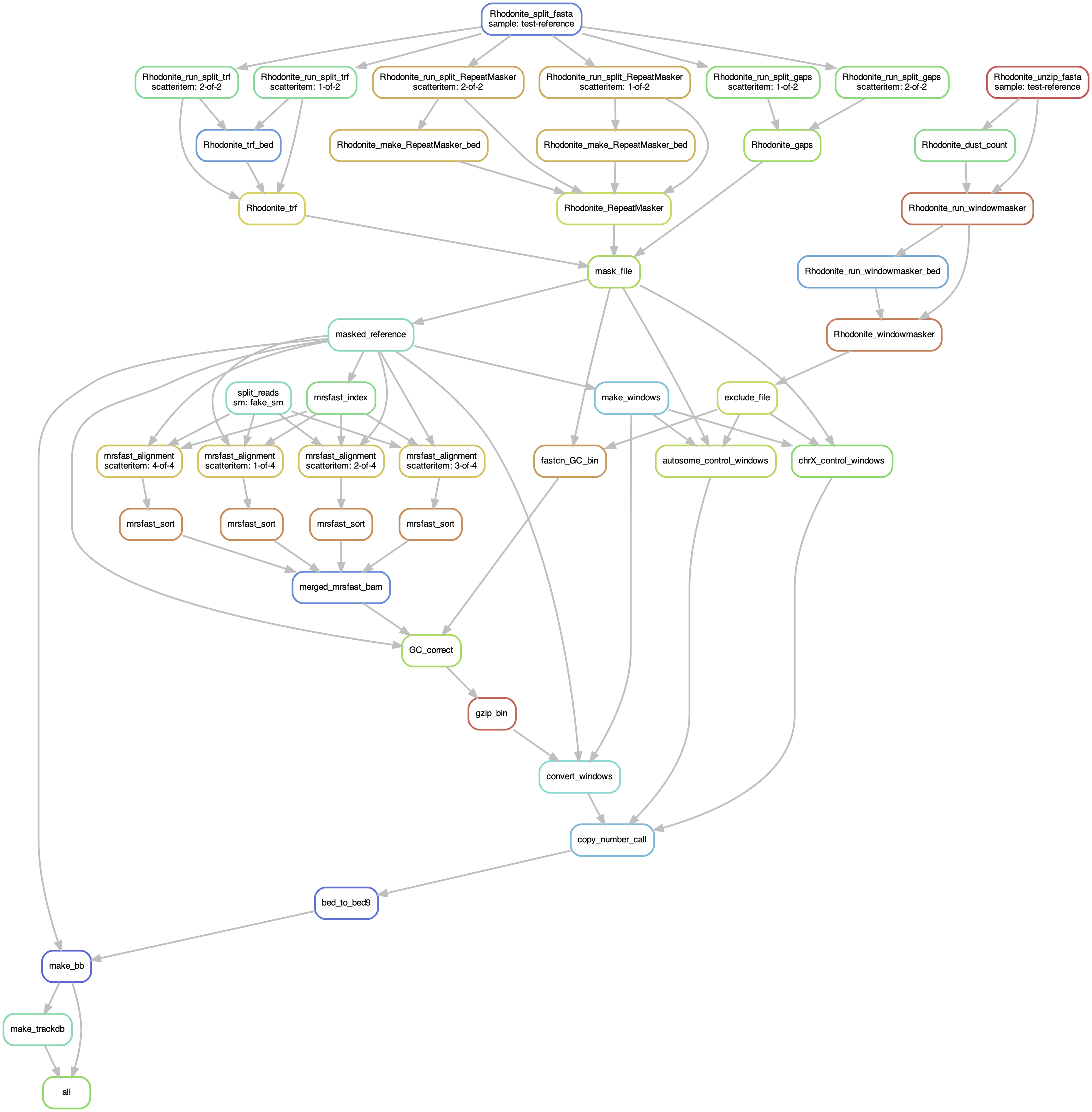
Workflows
hap-windows
Python★ 2
Extract haplotype-specific sequences centered on BED intervals from phased VCFs.